August 4, 2025
From July 8th to 11th, three Epigeners traveled to Bordeaux to take part in JOBIM 2025, a key conference for computational biology. Epigene Labs presented two posters and gave a live demo of its open-source tool, InMoose. Here’s a look at what we shared with the scientific community.
A few years ago, Epigene Labs made a bold move: simplifying its data science stack by migrating fully to Python – leaving behind the traditional R/Python dual setup used in most bioinformatics workflows. This decision meant porting essential R-based tools to Python, as no equivalents were available at the time – an ambitious data engineering challenge.
The effort paid off. This initiative led to InMoose, a Python-native library that now reimplements trusted tools for bulk transcriptomic analysis, including limma, edgeR, DESeq2, ComBat, and ComBat-Seq. With three peer-reviewed validations, InMoose proves that robust data engineering can improve reproducibility without compromising performance.
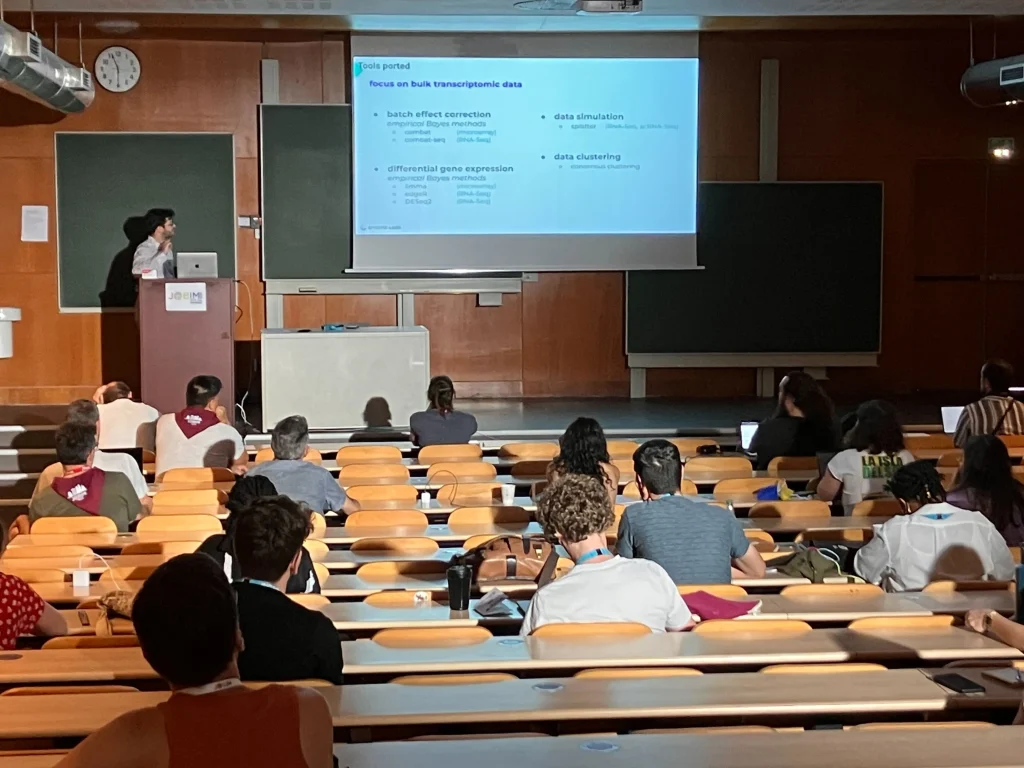
On July 8th, Maximilien Colange, lead developer of InMoose, walked attendees through the origins of the project and demonstrated how to use it effectively. Several presentations during the conference also featured research conducted using the InMoose environment, a strong sign of its growing adoption by the community.
Curious to try it yourself? You can access the tutorial as a shared notebook.

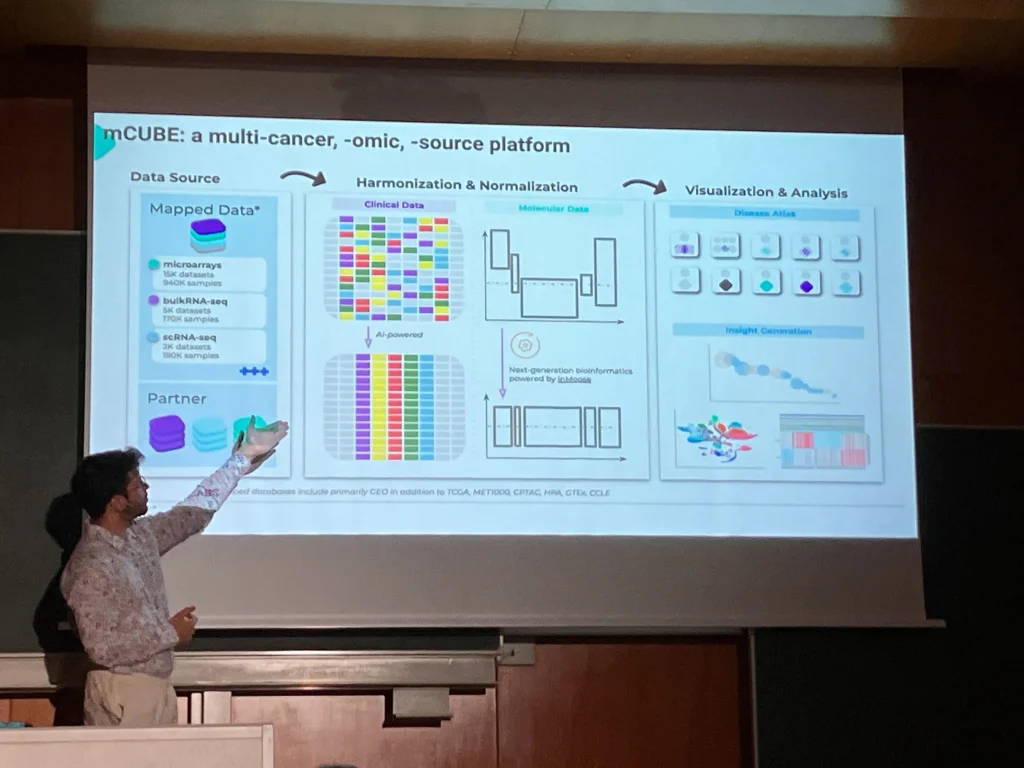
Robust scientific findings depend on reproducibility and generalizability. Larger cohorts are key to capturing biological variability and producing trustworthy results. Yet, omics data often suffers from fragmentation – different technologies, protocols, and naming conventions make datasets hard to combine, despite the clear value of doing so.
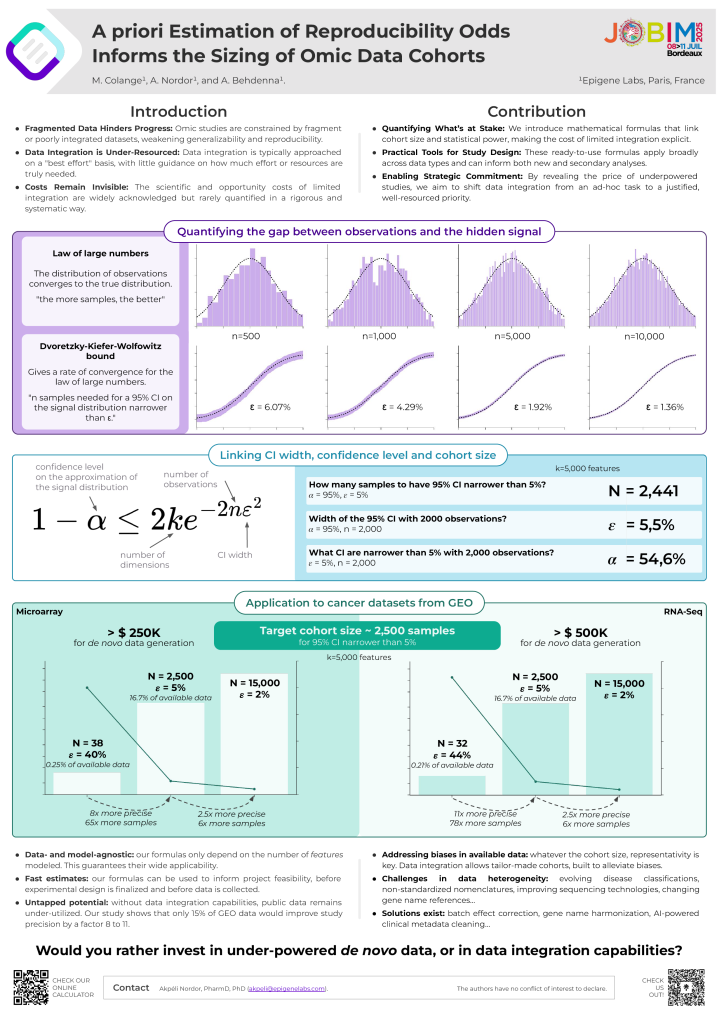
On July 10th, Maximilien Colange presented a methodology that helps quantify the trade-off between cohort size and analytical accuracy. This work provides a concrete answer to a long-standing question: How much value are we losing by working with small, fragmented datasets?
This work advocates for making data integration a strategic, well-funded data engineering priority, not an afterthought.
To support the community, we’ve also released an online calculator to help researchers assess the impact of their cohort size.
Antigen-targeting therapies have shown great promise in oncology – but most candidates still fail in clinical trials due to safety or effectiveness concerns. Identifying better targets earlier in the pipeline can dramatically improve success rates.
On July 10th, Léa Meunier, our senior computational biologist, presented Epigene’s scalable, data engineering-driven platform that leverages public omics datasets to discover and rank new antigen targets across various cancer types.
The pipeline, validated across 18 cancer types, uses a human-in-the-loop AI approach for data curation and harmonization (e.g., normalization, batch effect correction, gene name standardization), to integrate a comprehensive compendium of data: 8 protein knowledge databases, 5 bulk RNA-seq atlases, 20 tissue-specific single-cell RNA-seq datasets and 168 microarray studies assembled into cancer-specific cohorts with over 9,400 samples and 20,000 genes.
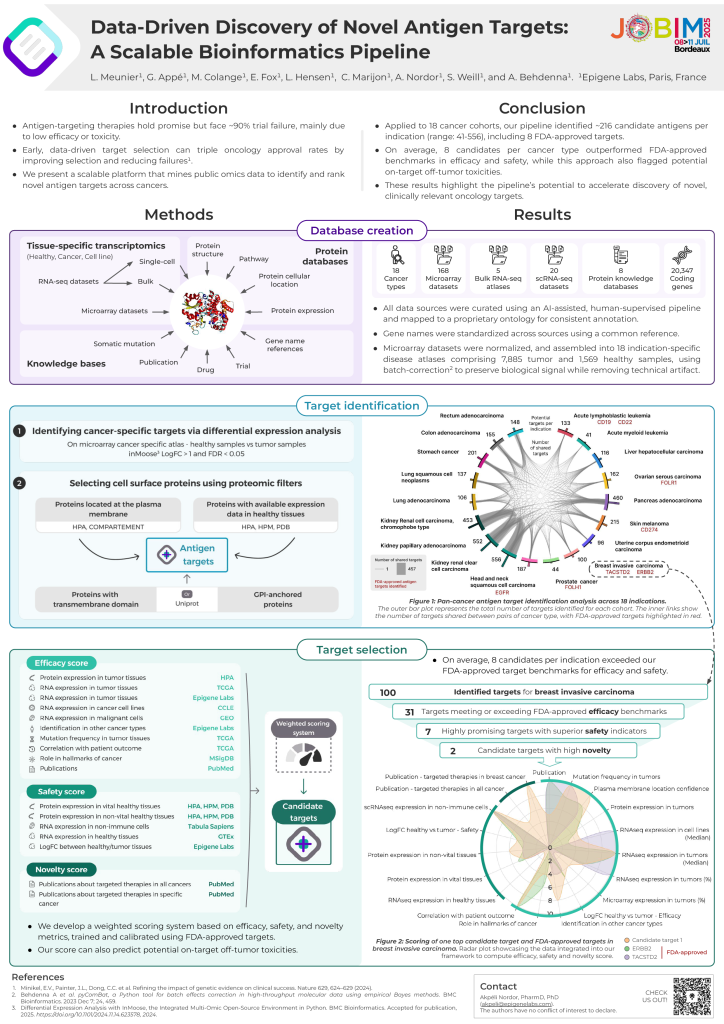

Key results:
This work highlights how data engineering in biopharma can accelerate and de-risk oncology drug development.
Authors: Maximilien Colange & Léa Meunier